The influenza A virus, which is responsible for seasonal flu outbreaks, is also the only influenza virus that has previously caused flu pandemics. This makes influenza A an important research topic, as the seasonal flu causes between 290,000 and 650,000 deaths per year globally. Because the influenza A virus is constantly changing, or mutating, it can be difficult to detect, treat, and inoculate against. To solve this problem, researchers are looking for parts of the influenza virus that do not change when the virus mutates. A panhandle structure on the virus known as the promoter region or promoter has emerged as a potential target.
In order to quickly detect the presence of the influenza A virus, researchers developed a fluorogenic probe that could bind to the promoter region of the influenza A virus RNA. A fluorogenic probe is based on small molecules called fluorophores that emit light when a specific target is present. In this study, the fluorogenic probe researchers created binds to part of the promoter region that consists of double-strand RNA structure carrying the internal loop, creating a significant light-up response that can identify the presence of influenza A.
The technique was presented in a paper published on May 23 in Analytical Chemistry.
"The promoter region of influenza A virus RNA has emerged as a new target for biochemical and therapeutic application because the sequences are not involved in the gene variations related to pathogenesis (how the flu virus develops) and antiviral resistance," said Yusuke Sato, an associate professor at Tohoku University. "These results represent the development of new molecular probes for influenza A research, with a view toward the diagnosis of influenza A infection, as well as the design of new antivirus drugs targeting the influenza A virus RNA promoter region."
In order to create the fluorogenic probe, researchers used a type of synthetic DNA called peptide nucleic acid (PNA). The triplex-forming PNA can be specifically developed to target the double-stranded RNA in the panhandle structure of the influenza A virus RNA in the sequence-selective manner. Researchers then combined the triplex forming PNA having a type of dye called thiazole orange with a small molecule that would bind with the internal loop structure of the RNA.
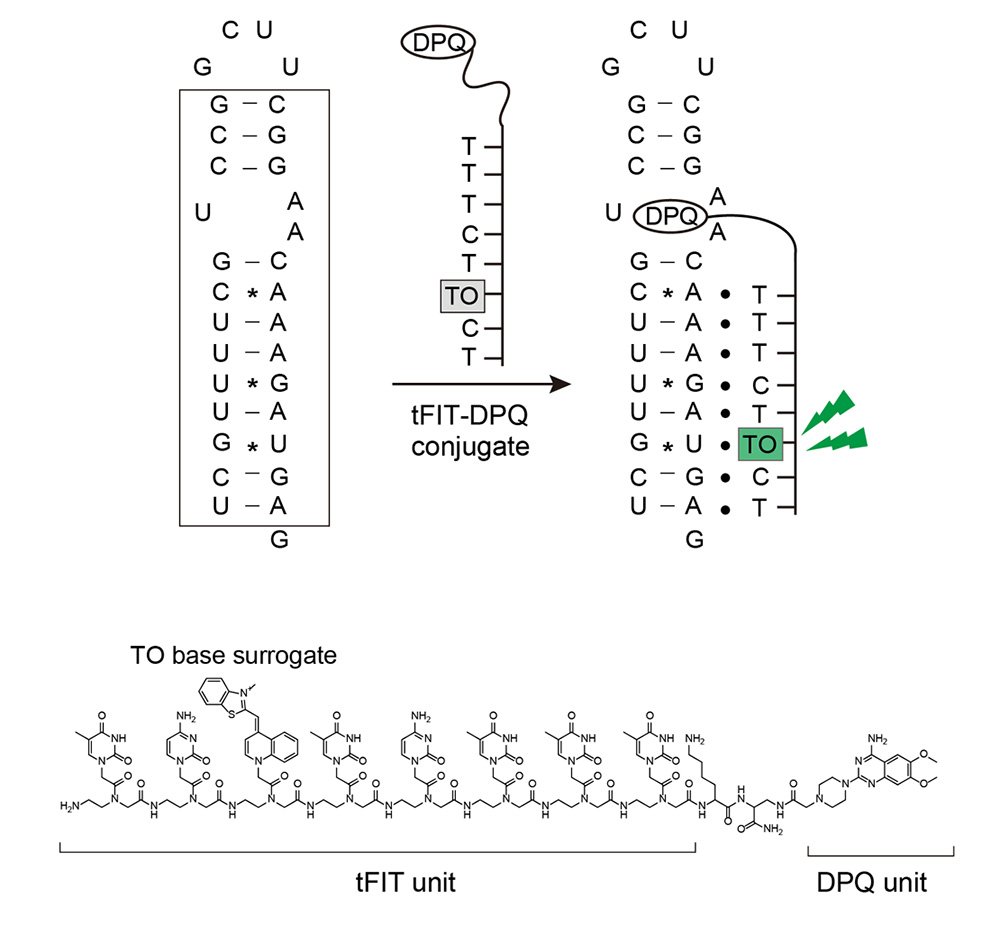
This combination is called a conjugate. To determine how effective the conjugate was, researchers first analyzed how brightly the conjugate glowed when it was bound to the target panhandle structure of the promoter region. It was more than 130-fold brighter than when it was not bound to anything. Compared to the small molecules alone, the combination of the PNA and the small molecules had a stronger binding affinity by two orders of magnitude. This result shows how promising this technique could be for the diagnosis of influenza A, since the promoter region remains stable no matter the strain of influenza.
"The research group demonstrated the selective fluorescence response of the conjugate for total RNA from influenza A virus H1N1-infected cells over that from mock-infected ones," said Sato. "This technique would serve as a promising candidate for the analysis of influenza A virus RNA based on the direct sensing of the influenza A virus RNA promoter region, in sharp contrast to the gold standard PCR method."
As the world keeps an eye on the ongoing COVID-19 pandemic, researchers are eager to find solutions now for future outbreaks of influenza A. By finding new ways to target specific parts of the influenza A virus that do not change when the virus mutates, this research could be used to create more sensitive tests that can detect the influenza A virus more easily. In the future, this could even be a promising target for antiviral drugs that could treat infections of influenza A.
- Publication Details:
Title: Fluorescence sensing of the panhandle structure of the influenza A virus RNA promoter by thiazole orange base surrogate-carrying PNA conjugated with small molecule
Authors: Yusuke Sato, Hiromasa Miura, Takaaki Tanabe, Chioma Uche Okeke, Kikuchi Akiko, Seiichi Nishizawa
Journal: Analytical Chemistry
DOI: 10.1021/acs.analchem.1c05488
Contact:
Yusuke Sato
Department of Chemistry, Graduate School of Science
Email: yusuke.sato.a7 tohoku.ac.jp
tohoku.ac.jp
Website: http://www.anal.chem.tohoku.ac.jp/index_en.html